 |
| Home | Research | Publications | Prospective Students | Teaching | Contact | Physics | Resources | Fun |
| Research Home
Current Projects Other Projects |
Novel computational approaches to predict misfolding-specific epitopes and design precision immunotherapeutics for neurodegenerative disease
We have developed a new way to rationally-predict disease-specific epitopes in misfolding prone proteins. I am co-inventor on >50 patents in the last 6 years on applications of this technology to personalized medicine in Alzheimer's disease (AD), Parkinson's disease (PD), and amyotrophic lateral sclerosis (ALS) immunotherapy, along with my experimental colleague Neil Cashman. The inventions focus on computationallypredicted diseasespecific epitopes in Amyloid beta and tau protein, two of the primary proteins whose misfolding and aggregation is implicated in AD. We have designed and developed conformationallyselective antibodies (Abs) that selectively bind these conformational epitopes. These precision Abs are designed to spare healthy protein, but target misfolded protein when it is behaving pathogenically. They are selective to toxic oligomers. We also pursue optimal epitopes and precision Abs for other proteins and diseases, including alphasynuclein in PD, and TDP43 in ALS. 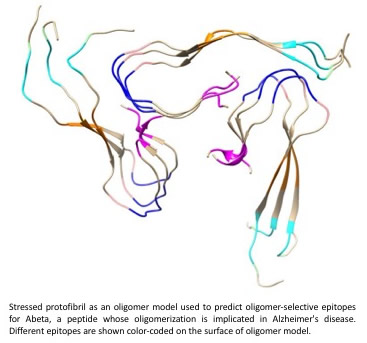
The methodology involves stressing structured fibrils computationally using a “collective coordinates” method (see this publication: PDF and also "Systems and Methods for Predicting Misfolded Protein Epitopes by Collective Coordinate Biasing" S.S. Plotkin, PCT/CA2016/051306).
Variations on irrational design are often used as a best guess for antibody development. I emphasize that rational design of antibodies from first principles using concepts from physical chemistry and structural molecular biology is a transformative concept in neurodegenerative disease immunotherapy, whose consequences are only beginning to be explored, and are being pioneered by our lab.
Given epitope predictions, we develop precision antibodies that best target diseased protein. Epitopes are optimally presented in cyclic peptides again using a computational design protocol which we have pioneered. Our Abs will ignore an epitope if the conformation is the same as in healthy protein, and our Abs are also designed to spare off-pathway targets in the human proteome. We use a multiplecriteria decision making (MDCM) scheme to screen and discover optimal precision Abs.
This work has been done in close collabo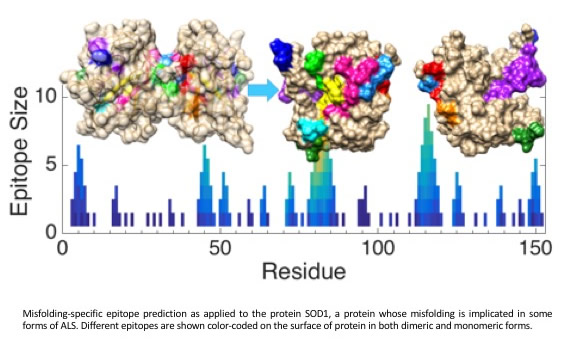 ration with my colleague Neil Cashman, in experimental neurology. The IP has been supported and licensed by ProMIS Neurosciences Inc., a development-stage biotech company developing precision therapeutics for the treatment of AD and ALS. I am the Chief Physics Officer for ProMIS Neurosciences (and maybe the only Chief Physics officer anywhere!)
ration with my colleague Neil Cashman, in experimental neurology. The IP has been supported and licensed by ProMIS Neurosciences Inc., a development-stage biotech company developing precision therapeutics for the treatment of AD and ALS. I am the Chief Physics Officer for ProMIS Neurosciences (and maybe the only Chief Physics officer anywhere!)
A novel feature of our AD antibodies directed against A-beta, which has resulted from our unique computational design approach, is that they do not bind Abeta plaques. This has been verified by immunohistochemistry of normal and AD patient brain sections. This is important, because most healthy but aged individuals have brains that are abundant with plaque, but they are free from AD symptoms because they do not carry toxic oligomeric strains of Abeta. The oligomer-selectivity of our antibodies is a unique strength: Many current commercial antibody hopefuls do in fact bind plaque, and as a result, treatment with these Abs may suffer from doselimitations related to the inflammation and edema they induce.
Designing Abs that are conformationally-selective to toxic oligomers is an extremely powerful technological approach, which has the potential to transform the way therapies are developed in the pharmaceutical industry, with profound implications for human health benefits of both Canadians and people abroad.
Related articles (see Publications):
Peng X, Cashman NR and Plotkin SS "Prediction of Misfolding-Specific Epitopes in SOD1 Using Collective Coordinates" J Phys Chem B, William A. Eaton Festschrift Special Issue, DOI: 10.1021/acs.jpcb.8b07680 (2018)
"Systems and Methods for Predicting Misfolded Protein Epitopes by Collective Coordinate Biasing" Inventor: Steven S. Plotkin, Provisional Patent Application # 62/253044, Filed November 9, 2015 (https://patentscope.wipo.int/search/en/detail.jsf?docId=WO2017079836)
McAlary L, Plotkin SS, Yerbury, J Cashman NR "Prion-like Propagation of Protein Misfolding and Aggregation in Amyotrophic Lateral Sclerosis" (in press) Frontiers Neuroscience (online at https://www.frontiersin.org/articles/10.3389/fnmol.2019.00262/full)
McAlary L, Plotkin SS, Cashman NR "Emerging developments in targeting proteotoxicity in neurodegenerative diseases" CNS Drugs (online at https://doi.org/10.1007/s40263-019-00657-9)
Gibbs E, Silverman JM, Zhao B, Peng X, Wang J, Wellington, CL, Mackenzie IR, Plotkin SS, Kaplan J, Cashman NR "A Rationally Designed Humanized Antibody Selective for Amyloid Beta Oligomers in Alzheimer’s Disease" Scientific Reports (2019) 9:9870 https://doi.org/10.1038/s41598-019-46306-5
Silverman JM, Gibbs E, Martens KM, Balducci C, Peng X, Wang J, Yousefi M, Cowan CM, Lamour G, Louadi A, Ban Y, Robert J, Stukas S, Forloni G, Hsiung GR, Plotkin SS, Wellington CL and Cashman NR "A Rational Structured Epitope Defines a Distinct Subclass of Toxic Amyloid-beta Oligomers" ACS Chem. Neurosci (2018)
"Epitopes in amyloid beta and conformationally-selective antibodies thereto" Inventors: Steven S Plotkin and Neil R Cashman, Provisional Patent 62/393,615 Filed Sept 14, 2016 (and related filings since 2016).
Home | Research | Publications | Prospective Students | Teaching | Contact | Physics | Resources || Fun |